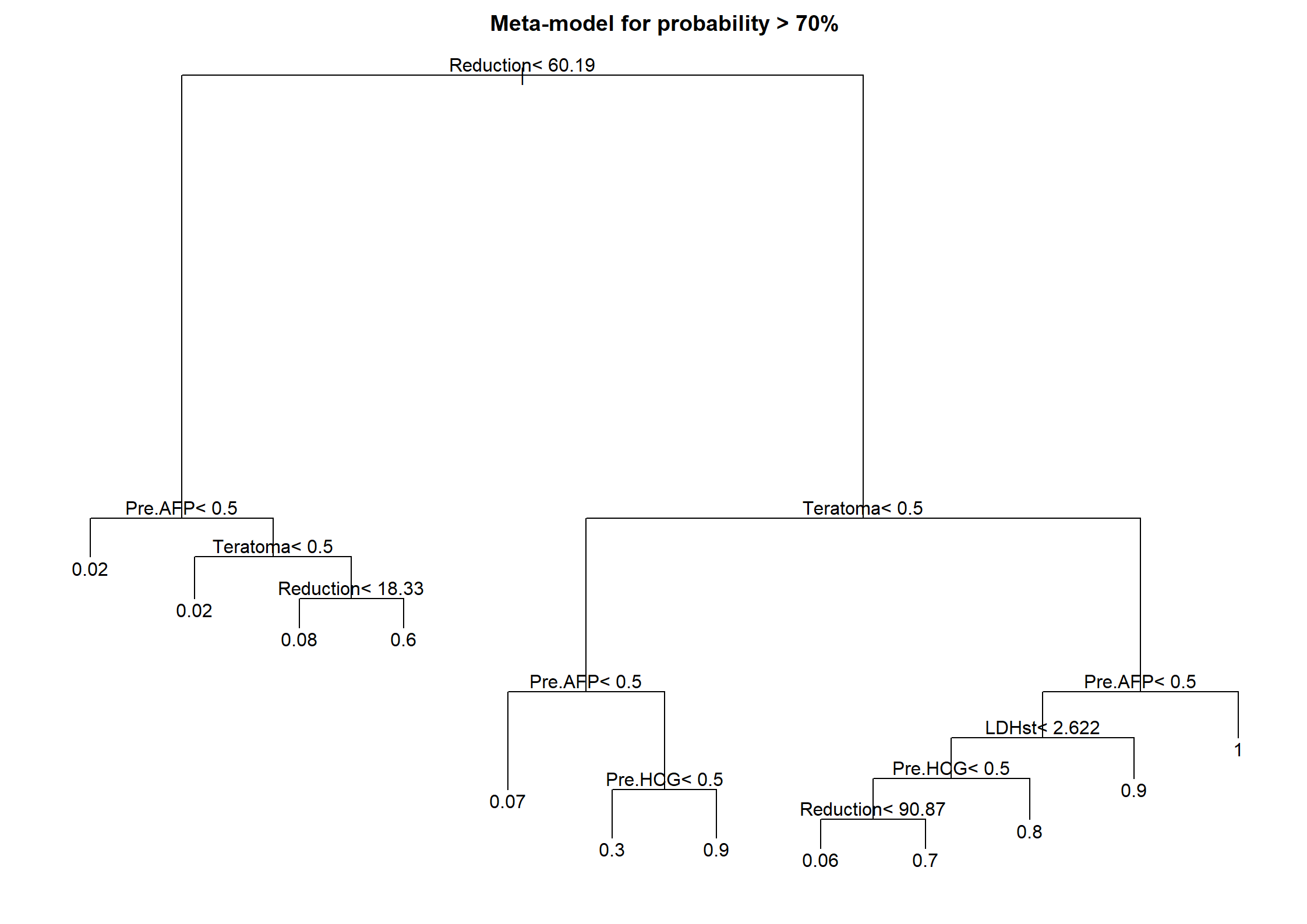
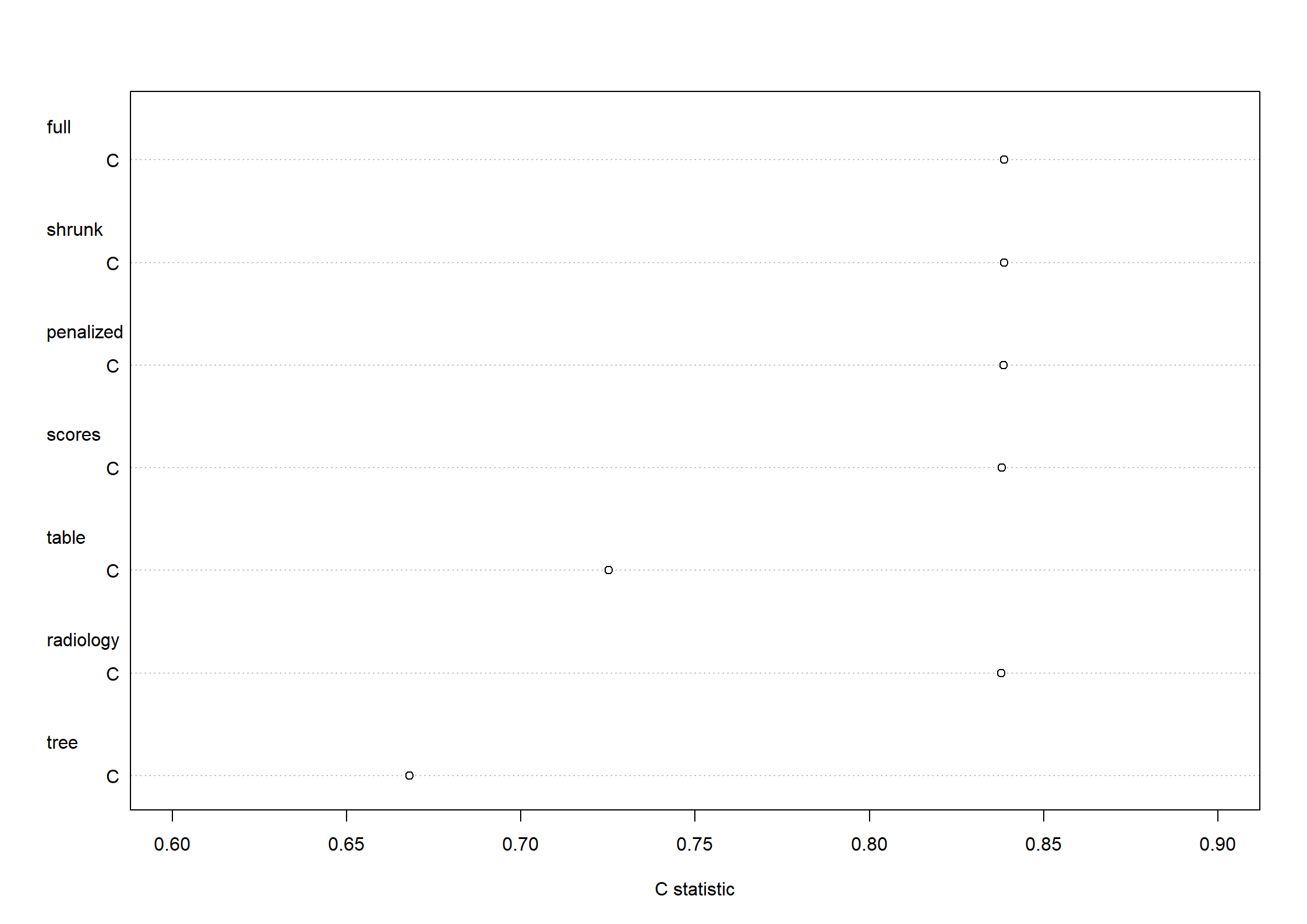
Code show/hide
# Fit a full 6 predictor model in n544
options(prType='html')
html(describe(n544), scroll=TRUE)11 Variables 544 Observations
Teratoma
| n | missing | distinct | Info | Sum | Mean |
|---|---|---|---|---|---|
| 544 | 0 | 2 | 0.746 | 252 | 0.4632 |
Pre.AFP
| n | missing | distinct | Info | Sum | Mean |
|---|---|---|---|---|---|
| 544 | 0 | 2 | 0.675 | 186 | 0.3419 |
Pre.HCG
| n | missing | distinct | Info | Sum | Mean |
|---|---|---|---|---|---|
| 544 | 0 | 2 | 0.704 | 205 | 0.3768 |
Pre.lnLDH
n missing distinct Info Mean pMedian Gmd .05 .10
544 0 472 1 0.4582 0.4191 0.7327 -0.38735 -0.26645
.25 .50 .75 .90 .95
-0.03266 0.31299 0.92886 1.33528 1.65328
| lowest : | -1.06582 | -0.948039 | -0.766913 | -0.731368 | -0.564972 |
| highest: | 2.42812 | 2.44938 | 2.48708 | 2.6237 | 2.76577 |
sqpost
n missing distinct Info Mean pMedian Gmd .05 .10
544 0 93 0.996 5.133 4.935 2.922 1.414 2.236
.25 .50 .75 .90 .95
3.162 4.472 6.652 9.669 10.000
lowest : 1.41421 1.73205 2 2.05477 2.23607 , highest: 10.9545 12.2474 13.0384 13.784 17.3205
reduc10
n missing distinct Info Mean pMedian Gmd .05 .10
544 0 200 0.998 4.482 4.833 4.307 -2.500 0.000
.25 .50 .75 .90 .95
2.000 5.167 7.337 8.785 10.000
lowest : -13.8095 -10 -8.33333 -8.10811 -6 , highest: 9.125 9.16667 9.28571 9.33333 10
Necrosis
| n | missing | distinct | Info | Sum | Mean |
|---|---|---|---|---|---|
| 544 | 0 | 2 | 0.743 | 245 | 0.4504 |
Tumor
| n | missing | distinct | Info | Sum | Mean |
|---|---|---|---|---|---|
| 544 | 0 | 2 | 0.743 | 299 | 0.5496 |
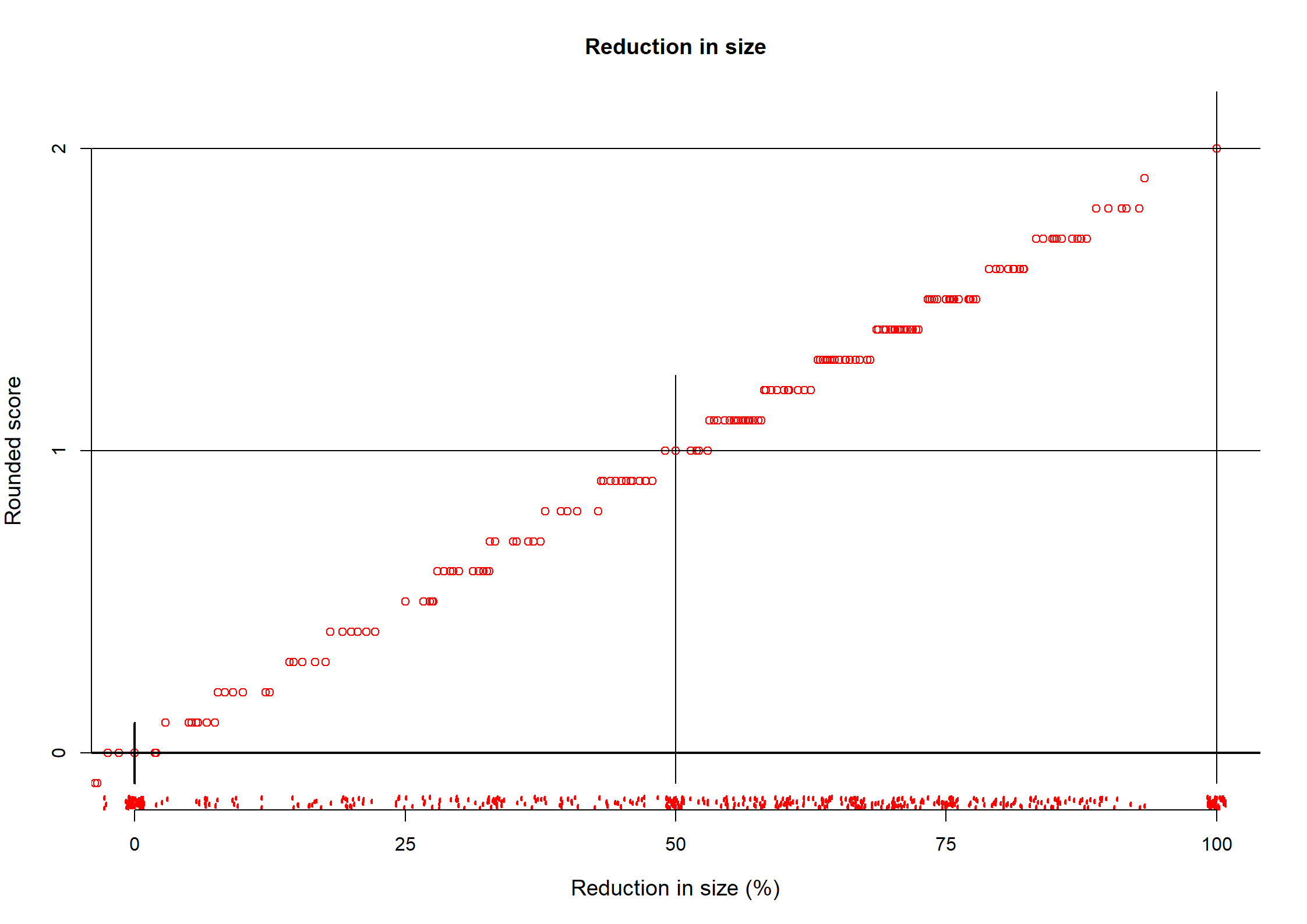
Reduction
n missing distinct Info Mean pMedian Gmd .05 .10
544 0 200 0.998 44.82 48.33 43.07 -25.00 0.00
.25 .50 .75 .90 .95
20.00 51.67 73.37 87.85 100.00
lowest : -138.095 -100 -83.3333 -81.0811 -60 , highest: 91.25 91.6667 92.8571 93.3333 100
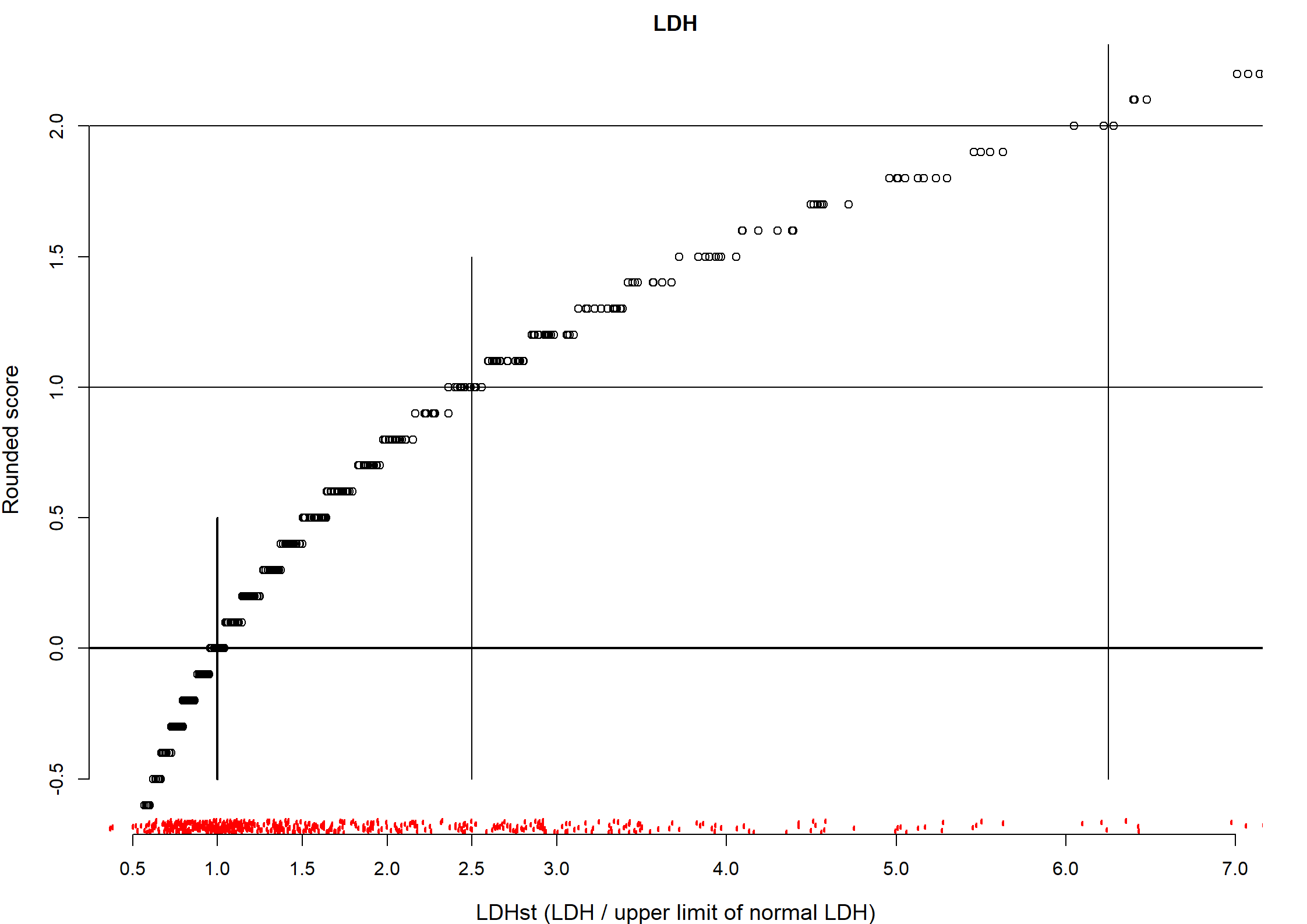
LDHst
| n | missing | distinct | Info | Mean | pMedian | Gmd | .05 | .10 | .25 | .50 | .75 | .90 | .95 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 544 | 0 | 472 | 1 | 2.03 | 1.693 | 1.605 | 0.6789 | 0.7661 | 0.9679 | 1.3675 | 2.5317 | 3.8014 | 5.2242 |
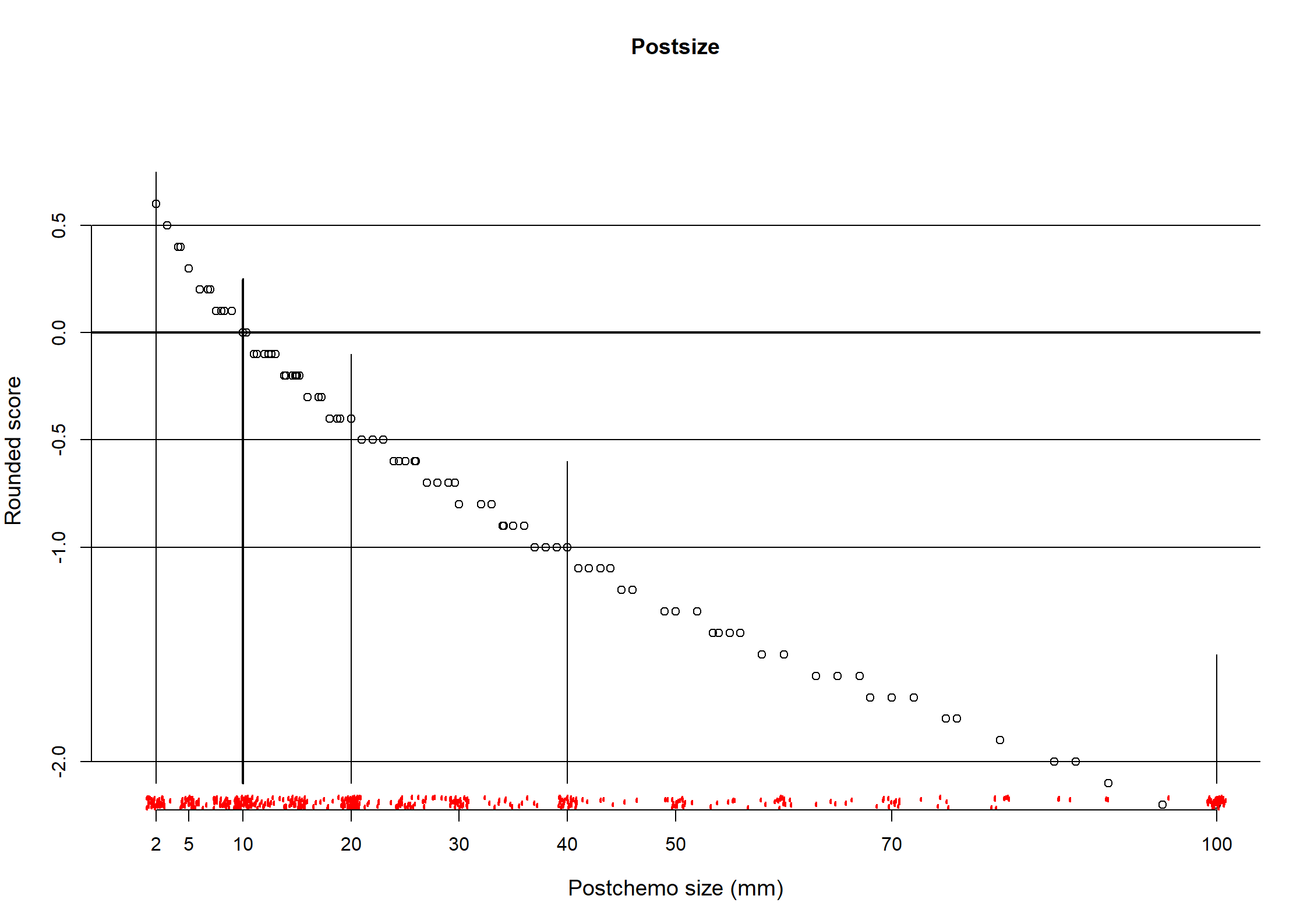
Post.size
n missing distinct Info Mean pMedian Gmd .05 .10
544 0 93 0.996 33.26 26.74 33.37 2.00 5.00
.25 .50 .75 .90 .95
10.00 20.00 44.25 93.50 100.00
lowest : 2 3 4 4.22209 5 , highest: 120 150 170 190 300
Code show/hide
# Start the fitting of 3 models
set.seed(1)
full <- lrm(Necrosis ~ Teratoma+Pre.AFP+Pre.HCG+log(LDHst)+sqrt(Post.size)+Reduction,
data=n544,x=T,y=T,linear.predictors=T)
print(full)Logistic Regression Model
lrm(formula = Necrosis ~ Teratoma + Pre.AFP + Pre.HCG + log(LDHst) +
sqrt(Post.size) + Reduction, data = n544, x = T, y = T, linear.predictors = T)
| Model Likelihood Ratio Test |
Discrimination Indexes |
Rank Discrim. Indexes |
|
|---|---|---|---|
| Obs 544 | LR χ2 211.56 | R2 0.431 | C 0.839 |
| 0 299 | d.f. 6 | R26,544 0.315 | Dxy 0.677 |
| 1 245 | Pr(>χ2) <0.0001 | R26,404 0.399 | γ 0.677 |
| max |∂log L/∂β| 1×10-7 | Brier 0.163 | τa 0.336 |
| β | S.E. | Wald Z | Pr(>|Z|) | |
|---|---|---|---|---|
| Intercept | -1.0425 | 0.6086 | -1.71 | 0.0867 |
| Teratoma | 0.9094 | 0.2140 | 4.25 | <0.0001 |
| Pre.AFP | 0.9025 | 0.2333 | 3.87 | 0.0001 |
| Pre.HCG | 0.7827 | 0.2305 | 3.40 | 0.0007 |
| LDHst | 0.9854 | 0.2089 | 4.72 | <0.0001 |
| Post.size | -0.2915 | 0.0815 | -3.58 | 0.0003 |
| Reduction | 0.0158 | 0.0052 | 3.04 | 0.0024 |