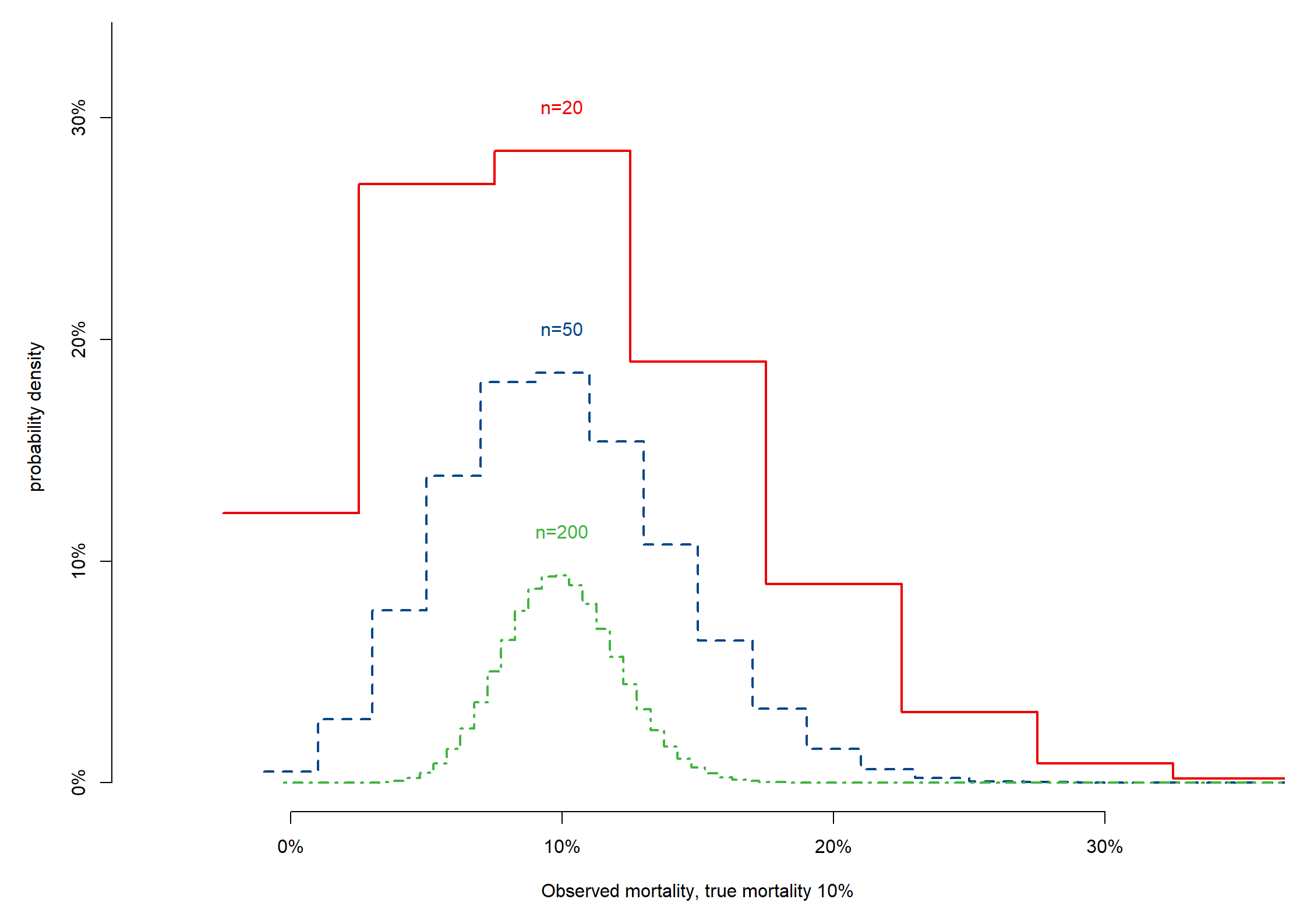
# Surg mortality; 10%
par(mfrow = c(1, 1), mar = c(5, 5, 1, 1))
for (mort in c(.1)) { ## ,0.05,0.02,.01)) { # 4 mortalities or only 1
plot(
x = seq(from = -.025, to = .975, by = .05), dbinom(x = 0:20, 20, mort), axes = F, type = "s", lwd = 2,
xlim = c(-.05, .35), ylim = c(0, .33), col = mycolors[2],
xlab = paste("Observed mortality, true mortality ", round(100 * mort, 0), "%", sep = ""), ylab = "probability density"
)
axis(side = 1, at = c(0, .1, .2, .3), labels = c("0%", "10%", "20%", "30%"))
axis(side = 2, at = c(0, 0.1, .2, .3, .4, .5, .6, .7), labels = c("0%", "10%", "20%", "30%", "40%", "50%", "60%", "70%"))
text(x = mort, y = .02 + dbinom(x = max(round(mort * 20), 0), 20, mort), labels = paste("n=20"), col = mycolors[2])
for (i in c(50, 200)) { # add more sample sizes
lines(
x = seq(from = 0 - (0.5 * 1 / i), to = 1 - (0.5 * 1 / i), by = 1 / i), dbinom(x = 0:i, i, mort),
type = "s", lty = ifelse(i == 50, 2, 4), lwd = 2, col = mycolors[ifelse(i == 50, 3, 4)]
)
text(x = mort, y = .02 + dbinom(x = max(round(mort * i), 0), i, mort), labels = paste("n=", i, sep = ""), col = mycolors[ifelse(i == 50, 3, 4)])
} # end loop n=50,200
} # end loop mort